Research Objects
Contents
Research Objects: The Braindump
David De Roure, July 2009
This is the myExperiment story on Research Objects. It's not the only take on this subject! The e-Laboratory groups has been meeting weekly, with representatives of myExperiment and many other projects, to build the bigger picture. There are important motivations coming from the clinical domain and also from the need to have a common object to exchange between the services that make up e-Laboratories. We don't address these here: this is Research Objects according to myExperiment.
From Workflows to Packs on myExperiment
The Web 2.0 Design Patterns tell us that "Data is the next Intel Inside", i.e.
Applications are increasingly data-driven. Therefore: For competitive advantage, seek to own a unique, hard-to-recreate source of data.
This reminds us that the value proposition of many Web 2 sites is their excellent support for one type of content. So we have photos on flickr, movies on YouTube and slides on SlideShare. And, when we began myExperiment, we became the unique, hard-to-recreate source of scientific workflows - specifically Taverna workflows. We are proud to hold the largest public collection of workflows available, and to have a growing number of workflow types, including Microsoft's Trident.
But, significantly, we also recognised that a workflow can be enriched as a sharable item by bundling it with some other pieces which make up the "experiment". We observe that researchers do not work with just one content type and moreover that their data is not in just one place - it's distributed, and sometimes quite messy too. So we have also developed support for “packs” – collections of items, both inside and outside myExperiment, which can be shared as one. Our user interface metaphor for packs is familiar from shopping sites - like a shopping cart or wishlist.
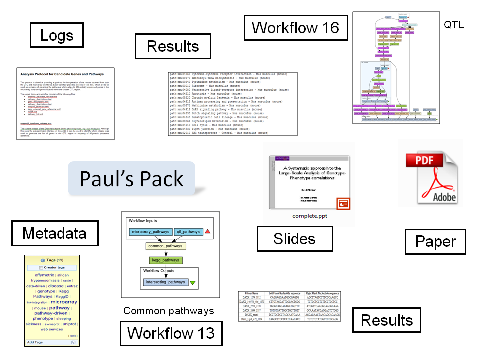
An excellent example of a pack is Pack 55, which is Paul Fisher's pack for his tryps work. It contains workflows, example input and output data, results, logs, PDFs of papers and slides - as depicted below.
Such a pack captures an experiment, it can be validated, it's self-described and it can be exported - we could say that packs "put the experiment into myExperiment"! The collection of packs is growing in variety and number. For example the pack of all the parts for a paper includes Paul's pack. There are packs of benchmark workflows, packs of example workflows, packs of papers and slides.
From Packs to Research Objects
As we have worked with our scientists and researchers we have studied the use cases for packs and watched how packs are actually being used. Through this we have recognised the emergence of a new form of digital object – the “Research Object”. Research objects are an evolution of packs and provide the sharable, reusable digital objects that enable research to be recorded and reused - which, fundamentally, is what Science and e-Research involves. In fact it seems to us entirely likely that, in the fullness of time, objects such as these will replace academic papers as the entities that researchers share, because they plug straight into the tooling of e-Research. It is Research Objects rather than papers that will be collected in our repositories. As well as a workflow repository, myExperiment has become a prototypical Research Object repository.
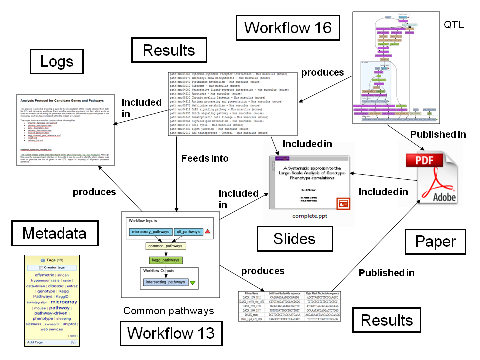
But what do these Research Objects look like? Paul annotated the pack with arrows and labels as shown below - this is a good example of what we need in a Research Object.
Also David Shotton in Oxford has also provided an excellent example of semantic publishing which comes at this from the other end - i.e. how do we augment a paper.
Our current pack user interface permits descriptions and comments so there is a way of expressing in free text the relationship of items to a pack. Some de facto practices are emerging, such as how presentations are described. One pack can be included in another. But currently we can't really express the relationships between things in packs, nor the relationships between packs themselves. Some of those labels will be domain-specific - we are implenting support for controlled vocabularies. Others may be more generic, such as basic notions of provenance.
We have a data model which includes workflows and packs and which we are extending to capture these relationships. Objects from myExperiment can be obtained in RDF from rdf.myexperiment.org. The data model uses a modularised ontology designed by David Newman which draws on Dublin Core, FOAF and also with Carl Lagoze's OAI Object Reuse and Exchange representation. We're engaged with Tim Clark's SWAN-SIOC work, and also with Luc Moreau and his Open Provenance Model (OPM) and Yolanda Gil's proposed W3C incubator group in provenance.
Meanwhile we are also gaining, from our users, a sense of what makes a good Research Object - for example, a workflow complete with example input and output data provides a means of checking the workflow still does what it used to (thanks to Werner Müller for highlighting this approach). One day perhaps we'll have a barometer telling you how good your Research Object is, like password security strength or filling out user profiles.
Defining Research Objects
To define a Research Object we need to understand its characteristics. We know that they are fundamentally about record and reuse - which means recording research for anticipated but also unanticipated re-use, and therein lines the challenge.
We've occasionally used the word "reproducible" in presentations, and there is a literature on "reproducible research". When I was working up my talk for the panel at ESWC 2009 I took the provocative position that a paper is just an "archaic human-readable form of a Research Object", and that in the future we won't say "can I have a copy of your paper please" but rather "could you share that research object with me please?" To present this I created a slide of words beginning with R (in fact, "Re.*") which characterise both future research and hence a Research Object, and this has evolved through several talks.
These are the Rs as they are now, on a Wiki page so they can continue to evolve in public. They have varied in number but settled down at six:
Replayable – go back and see what happened
Experiments are automated. They might happen in millseconds or in months. Either way, the ability to replay the experiment, and to study parts of it, is essential for human understanding of what happened.
Repeatable – run the experiment again
There's enough in a Research Object for the original researcher or others to be able to repeat the experiment, perhaps years later, in order to verify the results or validate the experimental environment. And the scale of data intensive science means lots of repetition of processing - to deal with the deluge of data or indeed the deluge of methods. Research Objects should help us scale.
Reproducible – run new experiment to reproduce the results
As every scientist knows, reproducible is different to repeatable! To reproduce (or replicate) a result is for someone else to start with the same materials and methods and see if a prior result can be confirmed.
Reusable – use as part of new experiments
One experiment may call upon another - an experiment may be used in a context other that that in which it was originally conceived. By assembling methods in this way we can conduct research, and ask research questions, at a higher level. Workflows are a great example.
Repurposeable – reuse the pieces in a new experiment
An experiment which is a black box is only reusaeable as a black box. By opening the lid we find parts, and combinations of parts, available for reuse, and the way they are assembled is a clue to how they can be re-used. This means there must be adequate description inside the box. Hence a research object is self-contained and self-describing - it contains enough metadata to have all the above characteristics and for a e-Lab component or service to make sense of it. (Computer Scientists have an "Re" word for this - "reflection", which means you can not only run a Research Object like a program but you can also look inside it like data).
Reliable – robust under automation
Automation brings systematic and unbiased processing, and also "unattended experiments" - human out the loop. In data-intensive science, Research Objects promote reliable experiments, but also they must be reliable for automated running.